When drug safety LLMs stop being demos and start becoming infrastructure
The most interesting thing in recent drug-safety LLM papers is not that the models are “good.” It is that the field is finally moving away from chatbot theater and toward workflow design. Across three recent papers, the same pattern shows up again and again: the winning systems are not the ones that sound the smartest. They are the ones that fit the task, fit the data, and fit the workflow.
That may sound obvious. But it matters, because pharmacovigilance is full of work that looks simple from far away and becomes messy the second you get close: free-text narratives, scanned documents, long labels, contradictory context, negation, uncertainty, duplicate concepts, and an endless need for traceability.
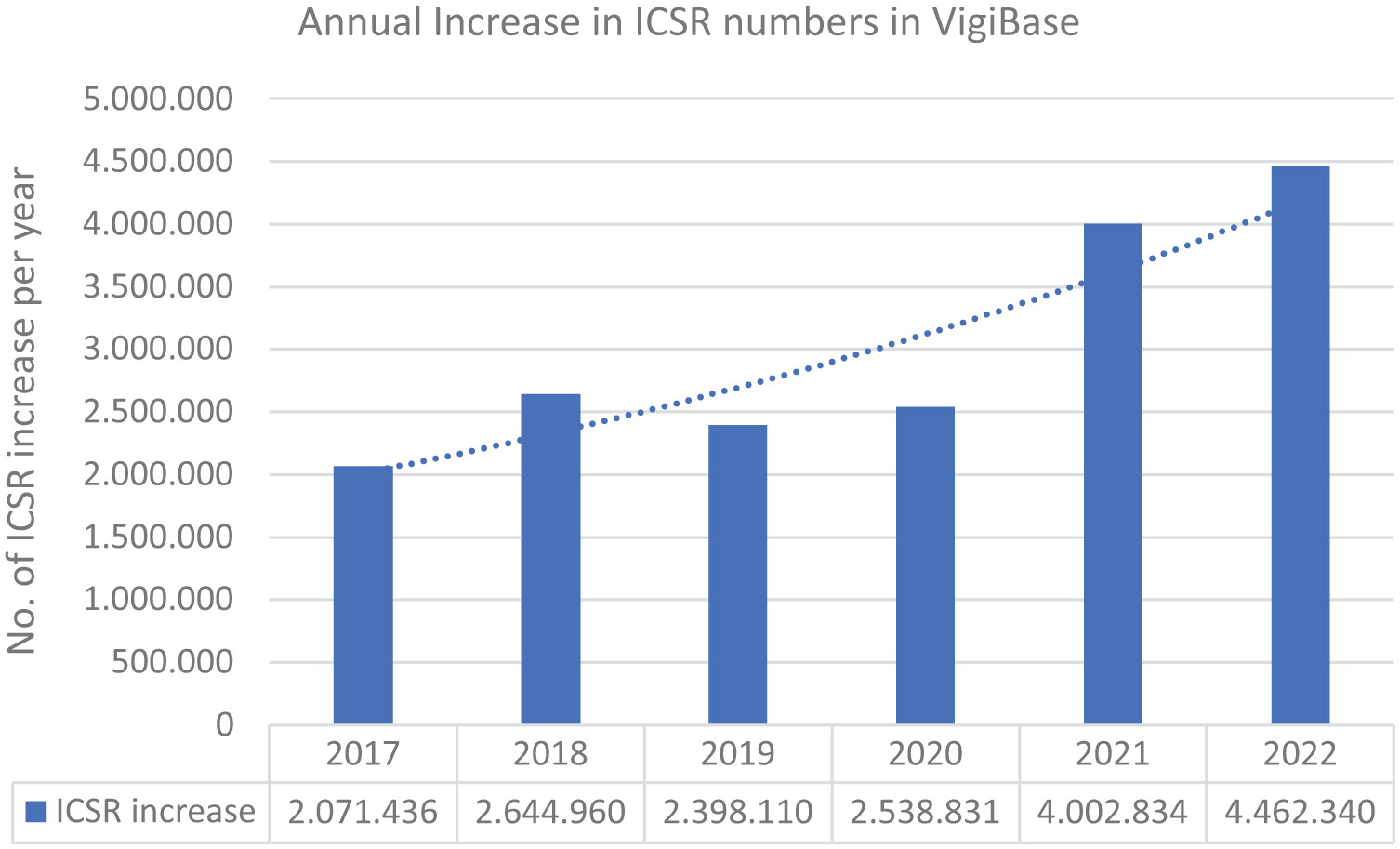
One paper frames the scale of the problem nicely.1 Annual increases in VigiBase Individual Case Safety Reports (ICSRs) rose from about 2.07 million in 2017 to 4.46 million in 2022, which helps explain why “just hire more reviewers” is not a serious long-term strategy anymore.
1. LLMs are most useful where the data are ugly
Roemming and colleagues focus on safety case intake, which is one of those operational areas where every inefficiency compounds downstream. Their proof-of-concept used LLMs to extract the minimum valid ICSR criteria plus additional safety-relevant fields from source documents. The headline result was not only decent extraction quality, but an estimated 39% efficiency gain, corresponding to about 20 minutes saved per case, with only 3.5% of extracted data points requiring manual correction.1
That is the kind of number that matters more than benchmark hype. It also explains why their five “points to consider” are so grounded: return on investment, early SME involvement, regulatory uncertainty, system integration, and organizational readiness. In other words, once an LLM leaves the demo stage, the real bottlenecks stop being model architecture and start being governance and process design.
This is probably the biggest shift in the literature right now. The question is no longer can a model extract a field? The question is can the extraction be trusted, reviewed, integrated, and defended in a regulated workflow?
2. In medication safety, context beats brute-force alerting
The second paper, by De Vito and colleagues, looks at medication safety from a different angle. Their HELIOT system uses clinical narratives plus pharmaceutical knowledge to decide whether an alert should be interruptive, non-interruptive, or suppressed. On a synthetic evaluation set, it achieved a macro F1 of 0.9869 and reduced interruptive alerts by 50.2% compared with traditional systems.2
That result matters because alert fatigue has been a chronic failure mode of clinical decision support for years. If a system warns about everything, clinicians learn to ignore it. The real goal is not maximum alert volume. It is maximum relevance. This is where LLMs have a genuine advantage: they can work with the narrative mess that older rules-based systems tend to miss. A patient’s prior tolerance, the wording of a previous reaction, the difference between allergy, intolerance, cross-reactivity, and vague historical suspicion — all of that often lives in text, not in neat database fields.
So the value proposition is not “LLMs replace clinical judgment.” It is that they can help surface the right warning at the right time, using context that older systems routinely throw away.
3. Bigger models are not automatically better models
The same paper also contains one of my favorite findings from the set: for direct drug-drug interaction prediction, smaller fine-tuned models outperformed larger ones. Their fine-tuned Phi-3.5 system reached average sensitivity 0.978 and accuracy 0.919 across 13 validation datasets, beating larger proprietary models and traditional baselines.2
That should make everyone in biomedical AI pause for a second.
We are so used to thinking in terms of model size that we sometimes forget the more important variables: task framing, adaptation strategy, input representation, and deployment constraints. In this case, the winning setup was not “use the biggest possible model.” It was “use a model that has been shaped for the exact problem you care about.”
For healthcare, that is encouraging. Smaller models are easier to run locally, easier to audit, cheaper to deploy, and better aligned with privacy-sensitive environments. That makes them much more plausible in actual hospitals and safety teams than a permanently remote, very large, general-purpose model.
4. The hard part is no longer extraction alone. It is extraction under context
The third paper, by Gisladottir and colleagues, gets at a subtle but important issue. They show that generative models like GPT-4 can match or even exceed prior task-specific systems for extracting adverse reactions from structured product labels, and can also generalize to extracting drug names from the drug-interaction section without additional fine-tuning.3
But their most interesting contribution is not just the headline performance. It is the breakdown of where extraction starts to fail. They found that performance changes depending on the label section, text length, and the linguistic complexity of the target concept. The “Adverse Reactions” section was easier than “Warnings and Precautions,” and longer texts tended to reduce precision. Most importantly, negated and discontinuous terms were much harder to capture reliably.3
A lot of biomedical NLP reporting still treats extraction as if it were a single task with a single score. But in practice, “extract the adverse reaction” means very different things depending on whether the concept is explicitly listed, buried inside warning language, hypothetically phrased, negated, or split across a sentence.
That is why these results feel important beyond SPL mining itself. They remind us that the true enemy is not low average performance. It is fragile performance in the difficult linguistic cases that matter most for safety interpretation.
So what is the actual takeaway?
Put the three papers together and a more mature picture emerges. LLMs look genuinely useful in drug safety when they are used as: workflow components, context readers, task-shaped systems ans human-review accelerators. That sounds less glamorous than “AI transforms pharmacovigilance,” but it is much more believable.
The real frontier is not whether an LLM can mention the right adverse event in a benchmark. It is whether a system can move safely across three layers at once:
- Language layer — messy narratives, labels, and source documents;
- Operational layer — time savings, alert burden, reviewer workload;
- Regulatory layer — traceability, validation, reproducibility, and defensibility.
The papers here suggest that progress is finally happening at all three layers together. The most interesting future for LLMs in pharmacovigilance is not the flashy assistant that writes elegant summaries. It is the quieter infrastructure underneath: extraction pipelines, triage support, alert prioritization, label mining, normalization, and human-in-the-loop review tools. Those systems are less visible. They are also the ones most likely to matter.
-
Roemming H-J, Hauben M, Wannhoff W, et al. How LLMs can advance safety case intake—points to consider and insights from a proof of concept. Therapeutic Advances in Drug Safety. 2025;16. DOI: 10.1177/20420986251386222. ↩ ↩2
-
De Vito G, Ferrucci F, Angelakis A. Enhancing Medication Safety with LLMs. Ital-IA 2025: 5th National Conference on Artificial Intelligence. 2025. ↩ ↩2
-
Gisladottir U, Zietz M, Kivelson S, et al. Leveraging Large Language Models in Extracting Drug Safety Information from Prescription Drug Labels. Drug Safety. 2025. DOI: 10.1007/s40264-025-01594-x. ↩ ↩2